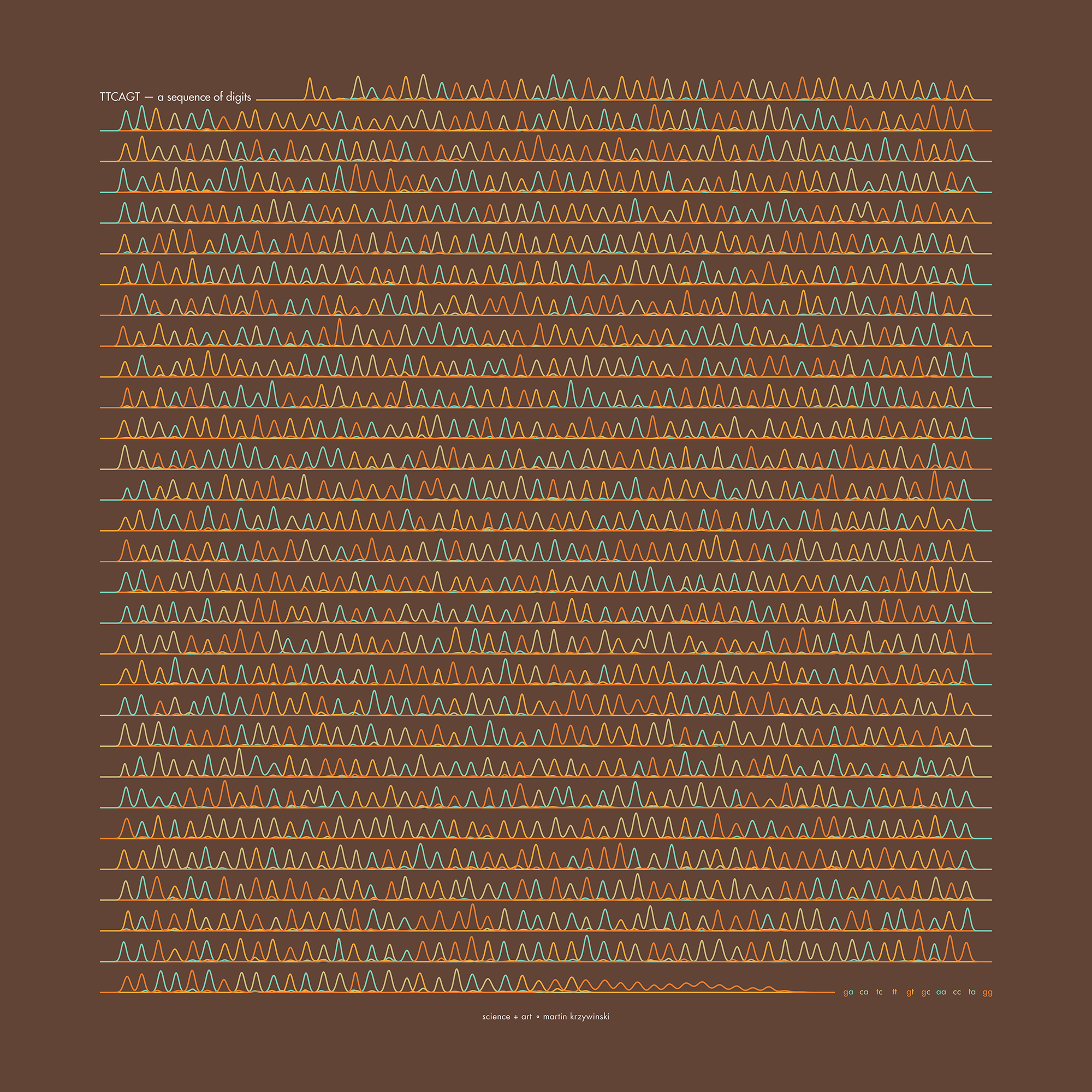 buy artwork
buy artwork
`\pi` Day 2014 Art Posters
On March 14th celebrate `\pi` Day. Hug `\pi`—find a way to do it.
For those who favour `\tau=2\pi` will have to postpone celebrations until July 26th. That's what you get for thinking that `\pi` is wrong. I sympathize with this position and have `\tau` day art too!
If you're not into details, you may opt to party on July 22nd, which is `\pi` approximation day (`\pi` ≈ 22/7). It's 20% more accurate that the official `\pi` day!
Finally, if you believe that `\pi = 3`, you should read why `\pi` is not equal to 3.
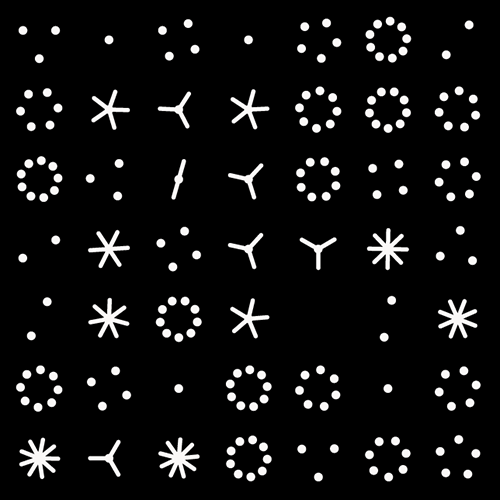
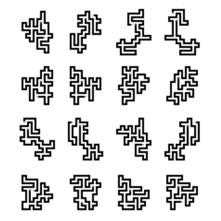
For the 2014 `\pi` day, two styles of posters are available: folded paths and frequency circles.
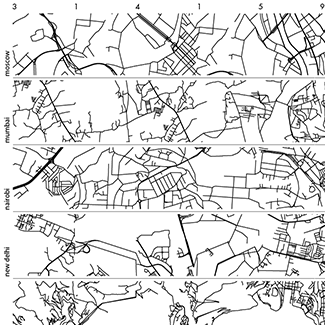
The folded paths show `\pi` on a path that maximizes adjacent prime digits and were created using a protein-folding algorithm.
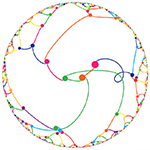
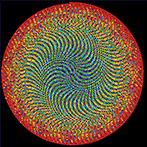
The frequency circles colourfully depict the ratio of digits in groupings of 3 or 6. Oh, look, there's the Feynman Point!
compute your own path
get simulation code
Download the HP lattice simulation binary. You'll need one of the three 2D methods — I used rem2dm, which does local and pull moves. If you'd like to learn more about the algorithm, read the publication.
A replica exchange Monte Carlo algorithm for protein folding in the HP model. Chris Thachuk, Alena Shmygelska and Holger H Hoos, BMC Bioinformatics 2007, 8:342 (17 Sep 2007).
download batch file
Download the batch file for 64- or 768-digit folding.
run simulation
When you run the 64-digit simulation, you're likely to find a path with E=-23, which is the lowest energy I've been able to sample. On my Intel Xeon E5540 (2.53 GHz) it takes anywhere from 1-30 seconds to find a E=-23 path (there are many possible paths at this energy), depending on the random seed. Here's the output of a typical run of the 64-digit folding simulation
> rem2dm -seq=hppphphphhhpphphhhppphpphhphhhphphppppphppphpphhhpphphpphpppphph
-maxT=220 -numLocalSteps=500 -eng=100 -maxRunTime=60 -traceFile=pi.64
-minT=160 -expID=pi.64 -numReps=10
REMC-HP2D-M
Begin Simulation
0.01: Current Best Solution: -8
0.01: Current Best Solution: -10
0.01: Current Best Solution: -13
0.02: Current Best Solution: -15
0.03: Current Best Solution: -16
0.03: Current Best Solution: -17
0.04: Current Best Solution: -18
0.04: Current Best Solution: -19
0.16: Current Best Solution: -20
0.27: Current Best Solution: -21
0.69: Current Best Solution: -22
36.23: Current Best Solution: -23
Real time: 120
ggslrrsrllssrrlrrllsrrlrrlslslrrsrlssrrsllrslrrlrsllsrsrrlsrssrs
p--h--p
| |
h--h h--p--p--p
| |
p--p h H h--p--p
| | | | |
p--h h--h--p p p--p
| | |
p--p--h h--p p--p p
| | | | |
h--h h h--p--h h--p
| | |
p--h h h--p--H h--p
| | | |
p--p p p--h--h
| |
p p--h--p
| |
p--p--h h
| |
p--p
End Simulation
If you want to apply this to different number (e.g.
φ
or
e
), you'll need to replace the digits with either p or h. Remember, the simulation will try to group the h's together. You can download 1,000,000 of
π
,
φ
and
e
.
The best path I could find for 768 digits is one with E=-223. In 1000s of simulations this solution came up only once. I also saw one path at E=-222. After that, there were many solutions at each of the less optimal energy levels.
If you manage to find a better one, let me know right away!
common problems
segmental fault
If you obtain a segmentation fault,
> ./rem2dlm REMC-HP2D-LM Begin Simulation Real time: 0 Segmentation fault
don't panic just yet. The folding binaries don't do a lot of error checking, so you have to get the input parameters correct.
For example, if you do not include the -eng parameter, the code will segfault.
Try one of the batch files above (64 digit batch file, 768 digit batch file) or the following simple job
> bin/rem2dm -seq=hhpppphhhhpppphh -maxRunTime=5 -eng 10
REMC-HP2D-M
Begin Simulation
3.13877e-17: Current Best Solution: -2
5.49284e-17: Current Best Solution: -3
1.0201e-16: Current Best Solution: -4
1.33398e-16: Current Best Solution: -5
Real time: 5
ggrllslsssrllsls
p--p--p
| |
h h--p
| |
H h
|
H h
| |
p--h h
| |
p--p--p
If this segfaults, then you'll need to recompile the code (see below).
compile code (optional—only if binaries don't work)
Precompiled binaries are available for download directly: rem2dm, rem2dlm, rem2dpm, rem3dm, rem3dlm, rem3dpm.
If these don't work on your system, you need to recompile them. Download the the protein folding code and see INSTALL.txt for compilation instructions.
Beyond Belief Campaign BRCA Art
Fuelled by philanthropy, findings into the workings of BRCA1 and BRCA2 genes have led to groundbreaking research and lifesaving innovations to care for families facing cancer.
This set of 100 one-of-a-kind prints explore the structure of these genes. Each artwork is unique — if you put them all together, you get the full sequence of the BRCA1 and BRCA2 proteins.
Propensity score weighting
The needs of the many outweigh the needs of the few. —Mr. Spock (Star Trek II)
This month, we explore a related and powerful technique to address bias: propensity score weighting (PSW), which applies weights to each subject instead of matching (or discarding) them.
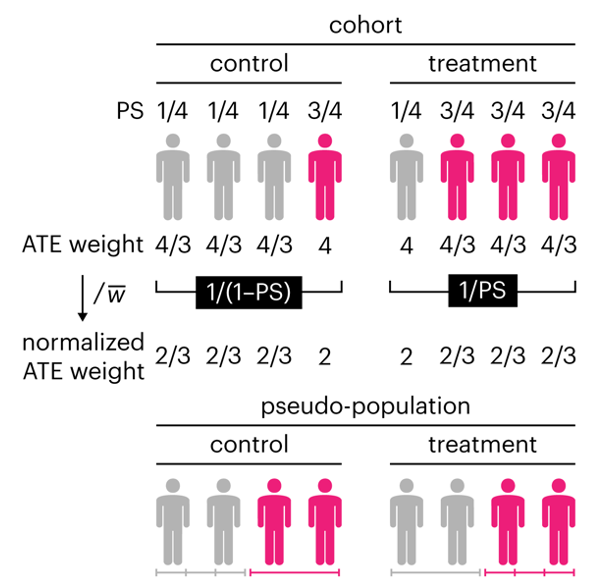
Kurz, C.F., Krzywinski, M. & Altman, N. (2025) Points of significance: Propensity score weighting. Nat. Methods 22:1–3.
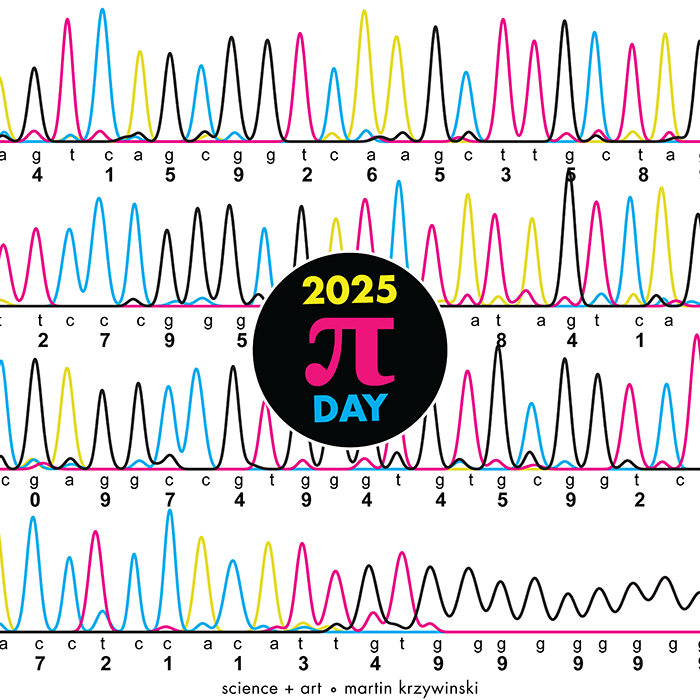
Happy 2025 π Day—
TTCAGT: a sequence of digits
Celebrate π Day (March 14th) and sequence digits like its 1999. Let's call some peaks.
Crafting 10 Years of Statistics Explanations: Points of Significance
I don’t have good luck in the match points. —Rafael Nadal, Spanish tennis player
Points of Significance is an ongoing series of short articles about statistics in Nature Methods that started in 2013. Its aim is to provide clear explanations of essential concepts in statistics for a nonspecialist audience. The articles favor heuristic explanations and make extensive use of simulated examples and graphical explanations, while maintaining mathematical rigor.
Topics range from basic, but often misunderstood, such as uncertainty and P-values, to relatively advanced, but often neglected, such as the error-in-variables problem and the curse of dimensionality. More recent articles have focused on timely topics such as modeling of epidemics, machine learning, and neural networks.
In this article, we discuss the evolution of topics and details behind some of the story arcs, our approach to crafting statistical explanations and narratives, and our use of figures and numerical simulations as props for building understanding.
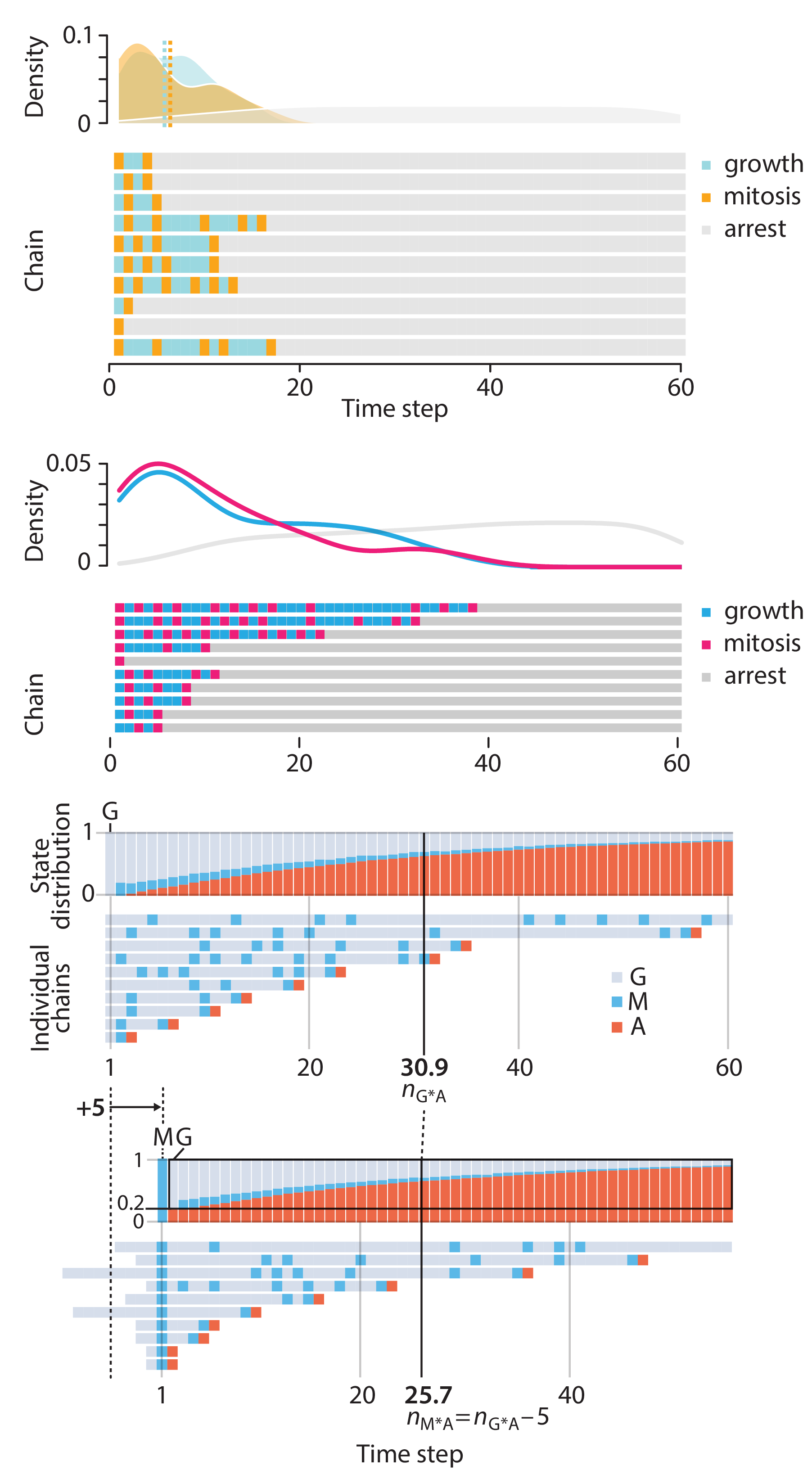
Altman, N. & Krzywinski, M. (2025) Crafting 10 Years of Statistics Explanations: Points of Significance. Annual Review of Statistics and Its Application 12:69–87.
Propensity score matching
I don’t have good luck in the match points. —Rafael Nadal, Spanish tennis player
In many experimental designs, we need to keep in mind the possibility of confounding variables, which may give rise to bias in the estimate of the treatment effect.
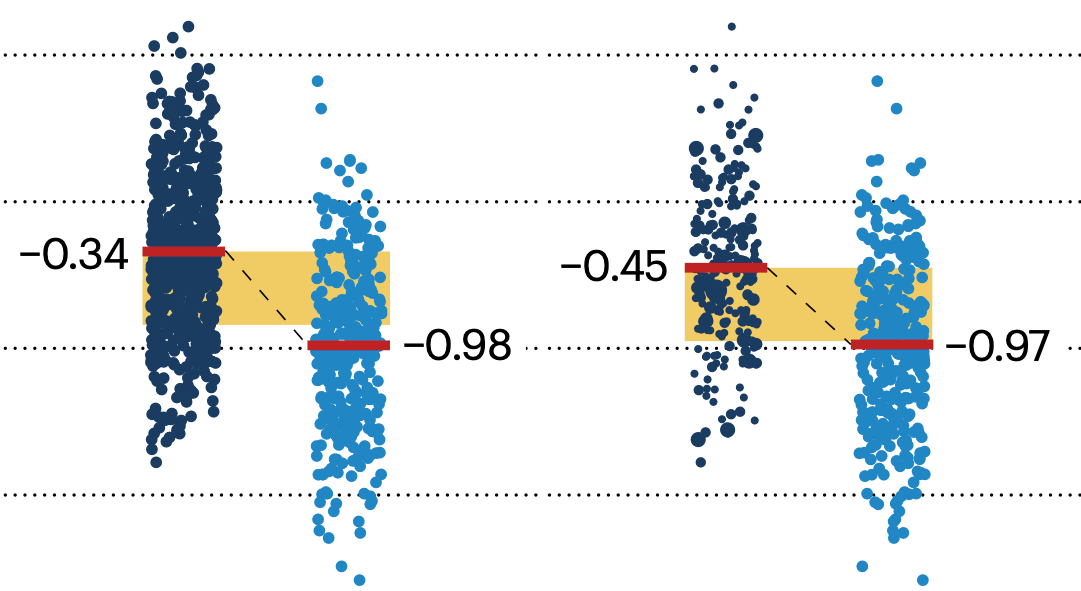
If the control and experimental groups aren't matched (or, roughly, similar enough), this bias can arise.
Sometimes this can be dealt with by randomizing, which on average can balance this effect out. When randomization is not possible, propensity score matching is an excellent strategy to match control and experimental groups.
Kurz, C.F., Krzywinski, M. & Altman, N. (2024) Points of significance: Propensity score matching. Nat. Methods 21:1770–1772.
Understanding p-values and significance
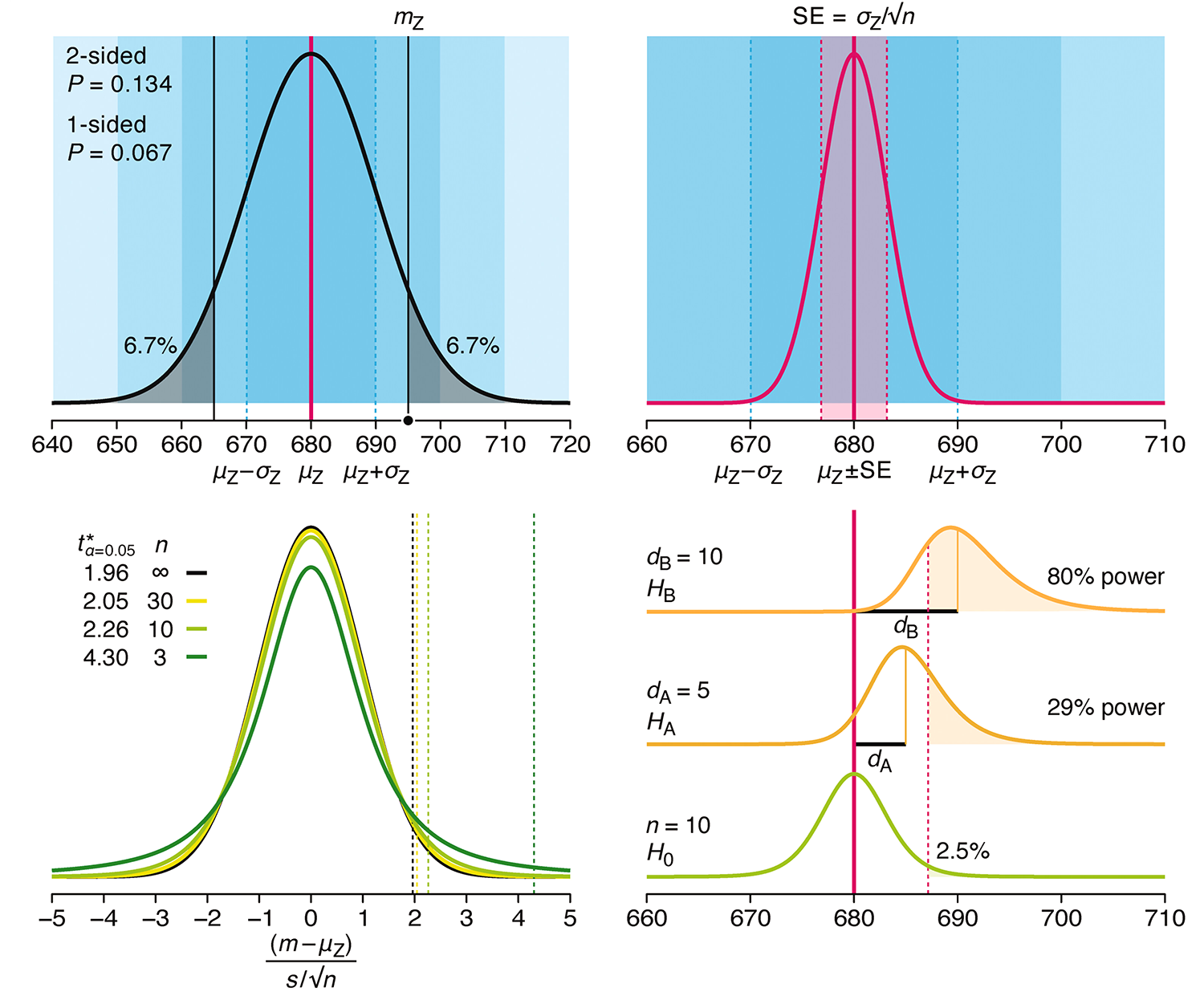
P-values combined with estimates of effect size are used to assess the importance of experimental results. However, their interpretation can be invalidated by selection bias when testing multiple hypotheses, fitting multiple models or even informally selecting results that seem interesting after observing the data.
We offer an introduction to principled uses of p-values (targeted at the non-specialist) and identify questionable practices to be avoided.
Altman, N. & Krzywinski, M. (2024) Understanding p-values and significance. Laboratory Animals 58:443–446.