Schemaball
Circular visualization of database schemas
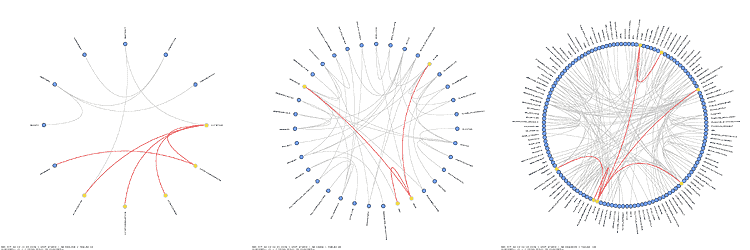
Schemaball was published in SysAdmin Magazine (Krzywinski, M. Schemaball: A New Spin on Database Visualization (2004) Sysadmin Magazine Vol 13 Issue 08). Who cites Schemaball?
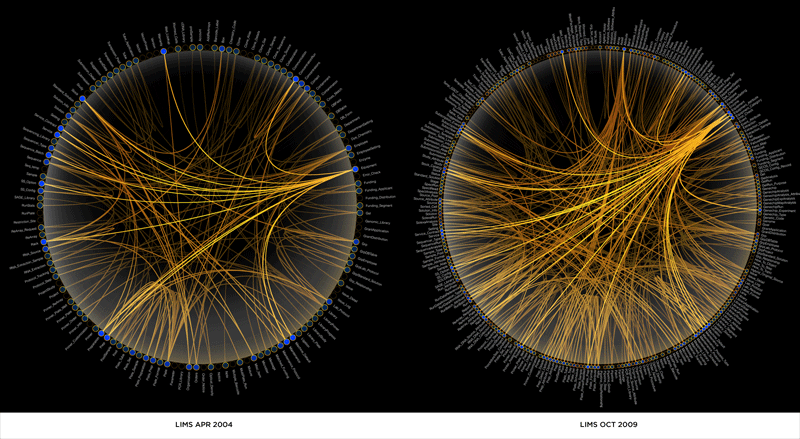
Schemaball is an SQL database schema viewer. It requires Perl and a few modules, such as GD and optionally Math::Bezier and SQL::Translator. Schemaball creates flexible visualizations of database schemas. Schemas may be read from an SQL schema dump, flat file or live database.
what is schemaball?
Schemaball is a flexible schema visualizer for SQL databases. The purpose of Schemaball is to help visualize the relationships between tables. Tables are related by foreign keys, which are fields which store the value of a record field from another table. Foreign keys create a lookup relationship between two tables. Large schemas can have hundreds of tables and table relationships. Keeping track of them call can be tedious, error-prone and slow down the schema development process. Schemaball provides a means to create flexible, static graphic images of a schema. Tables and table links can be hidden, highlighted and foreign key relationships can be traversed forward or backward to highlight connected tables.

Schemaball produces images called schema balls. Schema balls are schema visualizations in which tables are ordered along a circle with table relationships drawn as curves or straight lines. Using an input configuration file, all elements of the schema ball can be configured.
Schemaball is free software, licensed under GPL. It is written in Perl and requires a few CPAN modules to run. It's simple to use, while being able to produce high quality schema balls suitable for publication. In particular, you can use Schemaball to generate elements of a schema ball and then create a composite in an image editing program. One such result is shown in the figure at right.
A Need for Flexible Database Visualization
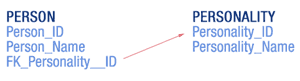

A database schema can be thought of as a directed graph. The nodes in the graph are the tables and the edges are relationships between them. The edges are created by foreign key fields in tables (see figure at right). The reason why the personality is stored in a separate table has to do with optimizing the way data is stored. There may be millions of records in the PERSON table, but only hundreds of different personalities. If the personality name is very long, the size of the database would be unnecessarily large since this same field value would be found in thousands of PERSON records. Storing a personality name in a separate table as a single record and associating it with a unique identifier, while other tables refer to it, is called normalization.
While in this example it's hard to get lost in the single table link, even with 10 tables keeping things straight can get messy. Schemaball helps to keep your head wrapped around your tables' connections. You can print out a schema ball and mark it up while you work on your schema. When I'm working on a database, I use the excellent mysqlfront to create tables and edit table fields. When I have my tables created, I print out a schema ball and mark up how I'd like to link the tables together. Once I create the foreign key fields, I can print out another schema ball and keep it for record.
Since it is designed primarily to Schemaball does not show the interal details of the tables. In future versions, support for display of fields may be included, to generate schema superballs.
Schemaball is at end of life
I am not longer actively working on Schemaball. As a prototype system, it works well to quickly figure out the extent of connectedness between tables and create some visually appealing images.
I've since been using the circular composition form in my Circos project, which applies this layout to visualizing genomes (Circos is particularly popular in the field of comparative genomes, in which relationships between two or more genomes are analyzed and displayed).
Case Study — Ensembl
Ensembl balls! See the structure of each Ensembl database, as drawn with Schemaball.
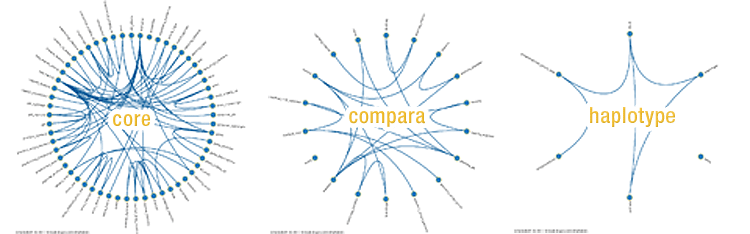
using Schemaball
To run Schemaball, you need Perl and a few CPAN modules. Schemaball has not been tested on Windows.
Propensity score weighting
It is not certain that everything is uncertain. —Blaise Pascal
We have already explored how we can mitigate bias caused by confounding variables in observational studies using propensity score (PS) matching (PSM) and propensity score weighting (PSW). However, any statistical model is only as good as its assumptions and, if it is specified incorrectly, it can itself produce biased estimates of the treatment effect.
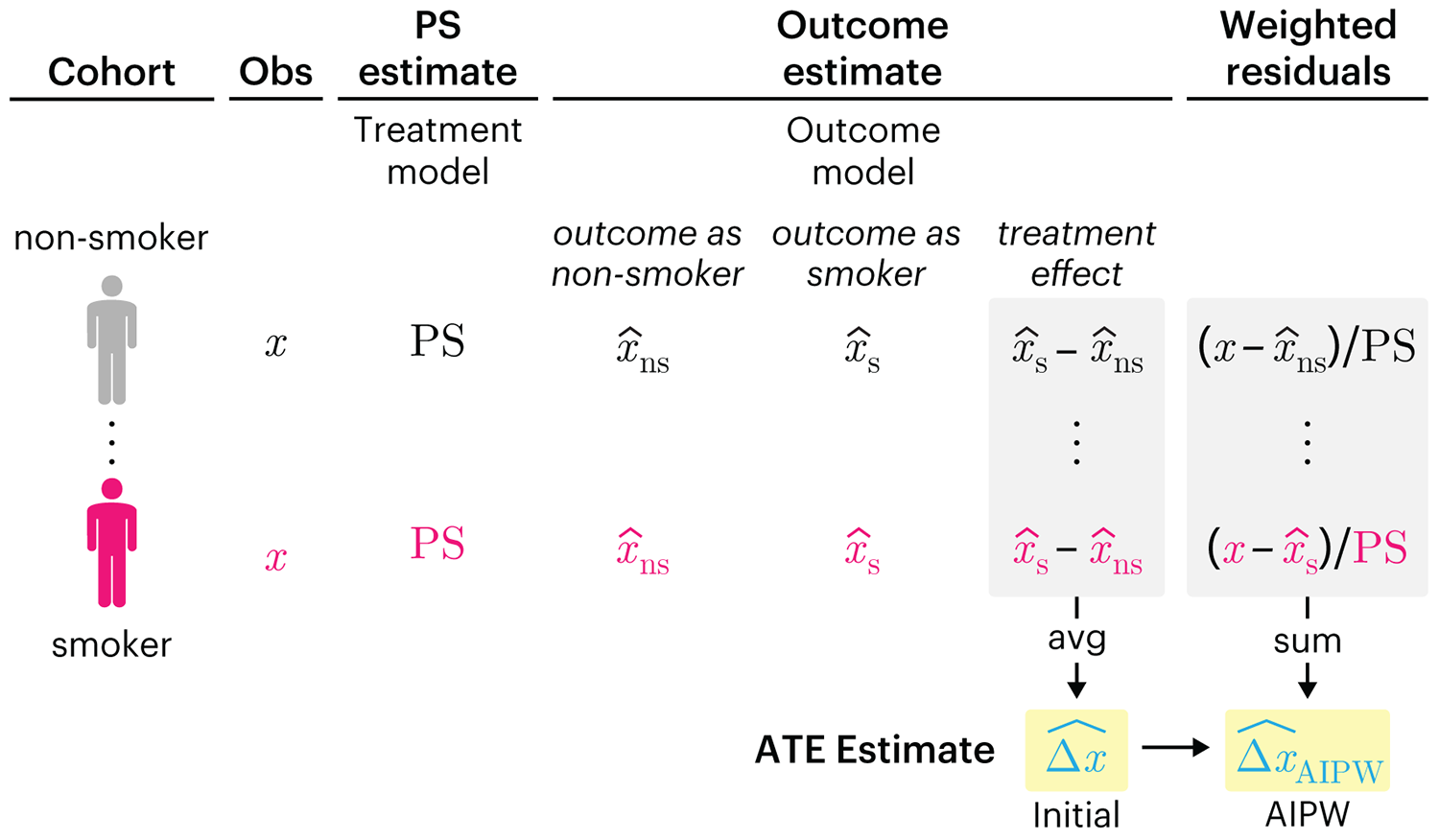
This month, we explore double robustness, a powerful statistical concept that provides a valuable “safety net” against the risk of an incorrect model. It offers two opportunities, instead of just one, to obtain a valid estimate of the treatment effect — making it possible to draw credible causal inferences from observational data without having to depend on a single set of modeling assumptions.
Kurz, C.F., Krzywinski, M. & Altman, N. (2026) Points of significance: Double Robustness. Nat. Methods 23:868–869.
Nature Biotechnology cover
My cover design on the 7 April 2026 Nature Biotechnology issue shows the dendrogram that represents a cluster of uniquely expressed (or downregulated) genes in human naive stem cells induced from such cells. Within each dendrogram block, the genomic barcode sequence (sampled from Supplementary Table 1) is depicted with a Code 39 barcode. The highlighted barcode is one of those used for cell isolation.
Ishiguro S. et al. A multi-kingdom genetic barcoding system for precise clone isolation (2026) Nature Biotechnology 44:616–629.

Browse my gallery of cover designs.

Happy 2026 π Day—
Art for the 5%
Celebrate π Day (March 14th) and enjoy the art — but only if you're part of the 5%.
Go ahead, see what you can't see.

Ishihara's Tests for Colour Deficiency
Authentic and accurate images of Ishihara's test plates photographed (and lovingly color-corrected) from the 38-plate Ishihara's Tests for Colour Deficiency.
I also provide the position, size, and color of each circle on each test plate.


Symmetric alternatives to the ordinary least squares regression
What immortal hand or eye, could frame thy fearful symmetry? — William Blake, "The Tyger"
This month, we look at symmetric regression, which, unlike simple linear regression, it is reversible — remaining unaltered when the variables are swapped.
Simple linear regression can summarize the linear relationship between two variables `X` and `Y` — for example, when `Y` is considered the response (dependent) and `X` the predictor (independent) variable.
However, there are times when we are not interested (or able) to distinguish between dependent and independent variables — either because they have the same importance or the same role. This is where symmetric regression can help.

Luca Greco, George Luta, Martin Krzywinski & Naomi Altman (2025) Points of significance: Symmetric alternatives to the ordinary least squares regression. Nat. Methods 22:1610–1612.