Learning Circos
Bioinformatics and Genome Analysis
Institut Pasteur Tunis Tunisia, December 10 – December 11, 2018
v2.00 6 Dec 2018 Download PDF slides
v2.00 6 Dec 2018
A 2- or 4-day practical mini-course in Circos, command-line parsing and scripting. This material is part of the Bioinformatics and Genome Analysis course held at the Institut Pasteur Tunis.
Quick links
BCGA 2018 | 1-day Circos course | Circos documentation best practices getting started | Brewer palette swatches | Color resources | Nature Methods Points of View Points of Significance
Visualizing and parsing .maf alignments
Additional material — Day 4
9h00 - 10h30 | Lecture 1 — Perl data structure and regular expression refresher
11h00 - 12h30 | Lecture (practical) 2 — Parsing .maf Ebola strain alignments on the command line
14h00 - 15h30 | Lecture (practical) 3 — Parsing .maf Ebola strain alignments with Perl
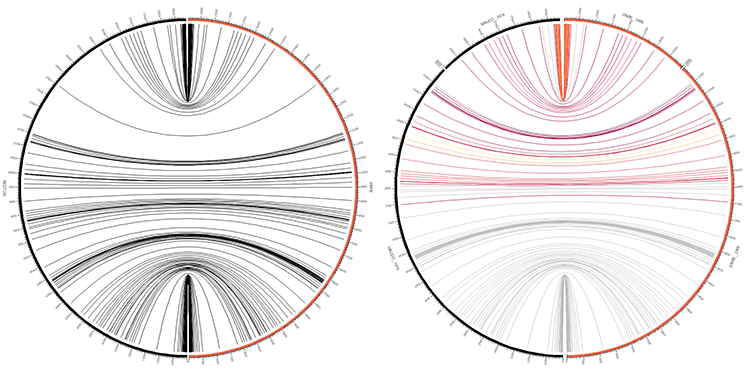
16h00 - 18h00 | Lecture (practical) 4 — Visualizing Ebola strain alignments
Concepts covered
.maf multiple-alignment data format, intermediate scripting, regular expressions, flexible parsing with Perl, creating Circos input data files, visualizing alignments between large number of sequences
A day's worth of additional material.